 | | PATHWAY: map00620 | |
| Entry |
|
|---|
| Name |
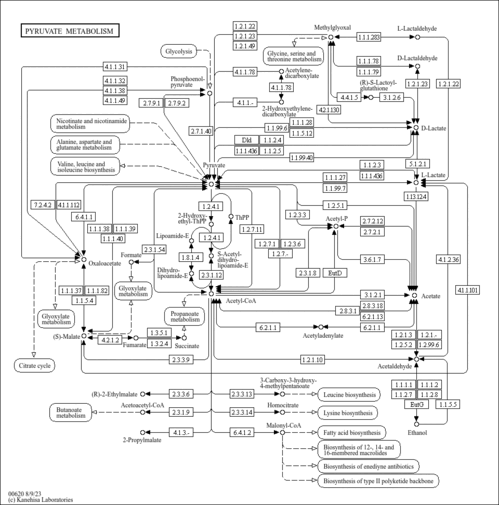
Pyruvate metabolism |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
|
|---|
| Module |
| M00172 | C4-dicarboxylic acid cycle, NADP - malic enzyme type [PATH:map00620] |
| M00579 | Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate [PATH:map00620] |
|
|---|
| Other DBs |
|
|---|
| Reference |
|
|---|
| Authors |
Nishizuka Y, Seyama Y, Ikai A, Ishimura Y, Kawaguchi A (eds). |
|---|
| Title |
[Cellular Functions and Metabolic Maps] (In Japanese) |
|---|
| Journal |
Tokyo Kagaku Dojin (1997) |
|---|
Related
pathway |
| map00250 | Alanine, aspartate and glutamate metabolism |
| map00260 | Glycine, serine and threonine metabolism |
| map00290 | Valine, leucine and isoleucine biosynthesis |
| map00522 | Biosynthesis of 12-, 14- and 16-membered macrolides |
| map00630 | Glyoxylate and dicarboxylate metabolism |
| map00760 | Nicotinate and nicotinamide metabolism |
| map01056 | Biosynthesis of type II polyketide backbone |
| map01059 | Biosynthesis of enediyne antibiotics |
|
|---|
| KO pathway |
|
|---|
|
|
|