| Entry |
|
|---|
| Name |
Atrazine degradation |
|---|
| Class |
Metabolism; Xenobiotics biodegradation and metabolism
|
|---|
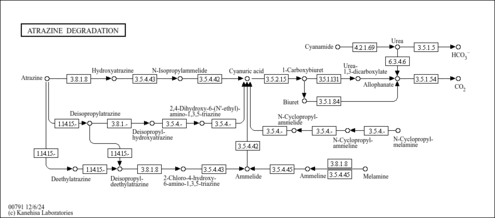
| Pathway map |
|
|---|
| Other DBs |
|
|---|
| Reference |
|
|---|
| Authors |
Shapir N, Cheng G, Sadowsky MJ, Wackett LP |
|---|
| Title |
Purification and characterization of TrzF: biuret hydrolysis by allophanate hydrolase supports growth. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Cheng G, Shapir N, Sadowsky MJ, Wackett LP. |
|---|
| Title |
Allophanate hydrolase, not urease, functions in bacterial cyanuric acid metabolism. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Li J, Biss M, Fu Y, Xu X, Moore SA, Xiao W |
|---|
| Title |
Two duplicated genes DDI2 and DDI3 in budding yeast encode a cyanamide hydratase and are induced by cyanamide. |
|---|
| Journal |
|
|---|
| KO pathway |
|
|---|