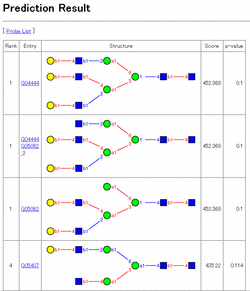
Example of prediction result
|
About N-glycan Prediction Server
Glycan biosynthesis is controlled by the expression of glycosyltransferases (GTs).
If the expression level of GTs is known in the transcriptome or proteome of a given organism, it is theoretically possible to predict the repertoire of glycan structures for an arbitrary set of experimental conditions (e.g., a particular disease or a particular tissue).
GECS is currently offered as a N-linked glycan (N-glycan) prediction tool, using (Affymetrix GeneChip) gene expression data to predict N-glycans. GECS works by automatically extracting the probe IDs and signal values of GTs, producing prediction scores for each N-glycan.
How to use
- Species
Users need to select a species. Currently, GeneChip datasets of mouse and human can be selected.
- Data Set
In the field labeled "Data Set 1", users provide a microarray expression data (text) file.
An example of gene expression data that can be used for input is available.
By inputting a second set of expression data, users can perform a comparative analysis of two sets of expression data. The details of this feature are explained below in the section "Operation".
- Server Option
Below each of the expression input boxes is a series of four buttons, labeled [1], [2], [1-2], and [2-1].
By selecting either [1] or [2], you can perform prediction of glycans for respective datasets 1 or 2.
By selecting [1-2], the difference in predicted score from Data Set 1 relative to the prediction score from Data Set 2 is shown (in other words, Score 1 - Score 2 is calculated).
By selecting [2-1], the opposite occurs, and the predicted difference is relative to the prediction score from Data Set 1 (Score 2 - Score 1 is calculated).
- Result Option
Users can specify the number of top ranking glycan predictions to display.
The amount of time to calculate and output results is affected by this value, so requesting an extremely large number of glycans to output will result in a longer time before a response is returned.
GECS can currently predict 2,466 glycans.
In the second input box, users can specify a threshold to indicate a special coloring (red) of those edges in the predicted glycan which have a signal value greater than or equal to the threshold.
Such edges will appear different than edges below the specified threshold (normally blue).
- Submit
After submitting your request, the server will require a few brief seconds to compute and rank predicted glycans before returning a response. It is not necessary to click the submit button repeatedly.
References
- Suga, A., Yamanishi, Y., Hashimoto, K., Goto, S., and Kanehisa, M.; An improved scoring scheme for predicting glycan structures from gene expressison data. Genome Informatics 18, 237-246 (2007). [pdf]
- Kawano, S., Hashimoto, K., Miyama, T., Goto, S., and Kanehisa, M.; Prediction of glycan structures from gene expression data based on glycosyltransferase reactions. Bioinformatics 21, 3976-3982 (2005). [pubmed]
Last updated: April 1, 2008
|
|

