KCF-S cluster No. 293 (23 metabolites)
Corresponding Phytochemical cluster No. 32
Plant Species
Cumulative plant class count
| class name | count |
|---|---|
| Liliopsida | 27 |
Cumulative family count
| class name | count |
|---|---|
| Zingiberaceae | 27 |
KEGG BRITE br08003

Categories (1)
| br08003 Category | # of metabolite |
|---|---|
| Zingiber derived compounds | 4 |
metabolites link (5)
Metabolite list (23)
| KNApSAcK ID | name | ChEMBL link | CTD link |
# of proteins in
ChEMBL interaction / related OMIM / related KEGG DISEASE |
# of genes in
CTD interaction / related diseases |
figure |
|---|---|---|---|---|---|---|
|
C00002746
|
[6]-Gingerdione
|
C050872
|

|
|||
|
C00002748
|
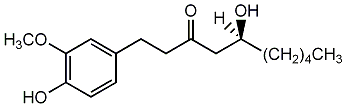
[6]-Gingerol
/ (S)-6-Gingerol / (S)-(+)-[6]Gingerol |
CHEMBL402978
CHEMBL1950582 |
4 / 4 / 0 |
|
||
|
C00002764
|
[6]-Paradol
|
CHEMBL2071440
|
C421614
|

|
||
|
C00002774
|
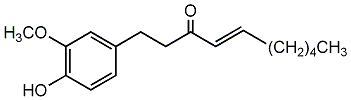
[6]-Shogaol
|
CHEMBL25948
|
C040115
|
8 / 3 / 4 | 10 / 1 |
|
|
C00031418
|
trans-10-Shogaol
|
CHEMBL24226
|
2 / 2 / 0 |

|
||
|
C00031419
|
trans-6-Shogaol
|
CHEMBL25948
|
8 / 3 / 4 |

|
||
|
C00031474
|
(S)-10-Gingerol
/ (+)-(S)-[10]-Gingerol |
CHEMBL549472
|
2 / 2 / 0 |

|
||
|
C00031475
|
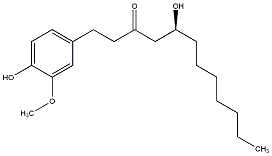
[8]-Gingerol
/ (S)-8-Gingerol / (+)-[8]-Gingerol |
CHEMBL1095671
|
2 / 2 / 0 |
|
||
|
C00034994
|
10-Shogaol
/ [10]-Shogaol |
CHEMBL24226
|
2 / 2 / 0 |

|
||
|
C00035032
|
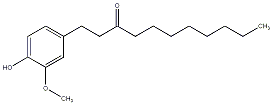
[7]-Paradol
|
CHEMBL431200
|
|
|||
|
C00035035
|
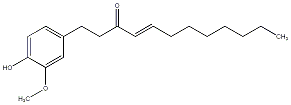
[8]-Shogaol
|
CHEMBL25893
|
2 / 2 / 0 |
|
||
|
C00035036
|
[9]-Paradol
|

|
||||
|
C00035442
|
[10]-Gingerdiol
|

|
||||
|
C00035443
|
[12]-Shogaol
|
CHEMBL24703
|

|
|||
|
C00035444
|
[4]-Gingerol
|

|
||||
|
C00035445
|
[4]-Isogingerol
|

|
||||
|
C00035446
|
[4]-Shogaol
|
CHEMBL277440
|

|
|||
|
C00035447
|
[6]-Gingerdiol
|
CHEMBL2071441
CHEMBL2071442 |

|
|||
|
C00035448
|
[7]-Gingerol
|

|
||||
|
C00035450
|
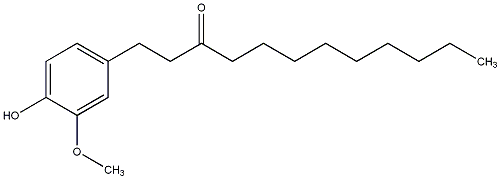
[8]-Paradol
|
|
||||
|
C00035687
|
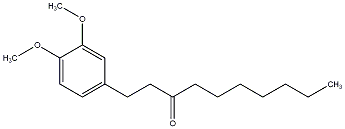
Methyl [6]-paradol
|
|
||||
|
C00035688
|
Methyl [6]-Shogaol
|

|
||||
|
C00035689
|
Methyl [8]-Shogaol
|

|
Human Protein / Gene in interactions
9 ChEMBL Protein in interactions
| accession | description | class description | KNApSAcK metabolite in interactions |
# of diseases
(OMIM / KEGG) |
|---|---|---|---|---|
| P08908 | 5-hydroxytryptamine receptor 1A | Serotonin receptor | C00002748 C00002774 C00031418 C00031419 C00031474 C00031475 C00034994 C00035035 | 1 / 0 |
| P08183 | Multidrug resistance protein 1 | drug | C00002748 C00002774 C00031418 C00031419 C00031474 C00031475 C00034994 C00035035 | 1 / 0 |
| Q8NER1 | Transient receptor potential cation channel subfamily V member 1 | TRPV (Vanilloid) | C00002748 C00002774 C00031419 | 0 / 0 |
| P10635 | Cytochrome P450 2D6 | Cytochrome P450 2D6 | C00002774 C00031419 | 1 / 0 |
| P35354 | Prostaglandin G/H synthase 2 | Oxidoreductase | C00002774 C00031419 | 0 / 3 |
| P08684 | Cytochrome P450 3A4 | Cytochrome P450 3A4 | C00002774 C00031419 | 0 / 1 |
| P09917 | Arachidonate 5-lipoxygenase | Oxidoreductase | C00002774 C00031419 | 0 / 0 |
| P23219 | Prostaglandin G/H synthase 1 | Oxidoreductase | C00002774 C00031419 | 0 / 0 |
| P49798 | Regulator of G-protein signaling 4 | Unclassified protein | C00002748 | 2 / 0 |
10 Gene in CTD interactions
| gene | gene name | gene description | KNApSAcK metabolite in interactions |
|---|---|---|---|
| 581 | BAX, BCL2L4 | BCL2-associated X protein |
C00002774
|
| 596 | BCL2, Bcl-2, PPP1R50 | B-cell CLL/lymphoma 2 |
C00002774
|
| 598 | BCL2L1, BCL-XL/S, BCL2L, BCLX, BCLXL, BCLXS, Bcl-X, PPP1R52, bcl-xL, bcl-xS | BCL2-like 1 |
C00002774
|
| 836 | CASP3, CPP32, CPP32B, SCA-1 | caspase 3, apoptosis-related cysteine peptidase (EC:3.4.22.56) |
C00002774
|
| 842 | CASP9, APAF-3, APAF3, ICE-LAP6, MCH6, PPP1R56 | caspase 9, apoptosis-related cysteine peptidase (EC:3.4.22.62) |
C00002774
|
| 1649 | DDIT3, CEBPZ, CHOP, CHOP-10, CHOP10, GADD153 | DNA-damage-inducible transcript 3 |
C00002774
|
| 1676 | DFFA, DFF-45, DFF1, ICAD | DNA fragmentation factor, 45kDa, alpha polypeptide |
C00002774
|
| 355 | FAS, ALPS1A, APO-1, APT1, CD95, FAS1, FASTM, TNFRSF6 | Fas cell surface death receptor |
C00002774
|
| 356 | FASLG, ALPS1B, APT1LG1, APTL, CD178, CD95-L, CD95L, FASL, TNFSF6 | Fas ligand (TNF superfamily, member 6) |
C00002774
|
| 142 | PARP1, ADPRT, ADPRT_1, ADPRT1, ARTD1, PARP, PARP-1, PPOL, pADPRT-1 | poly (ADP-ribose) polymerase 1 (EC:2.4.2.30) |
C00002774
|