| Entry |
|
|---|
| Name |
Oxidative phosphorylation - Janthinobacterium sp. Marseille |
|---|
| Class |
Metabolism; Energy metabolism
|
|---|
| Pathway map |
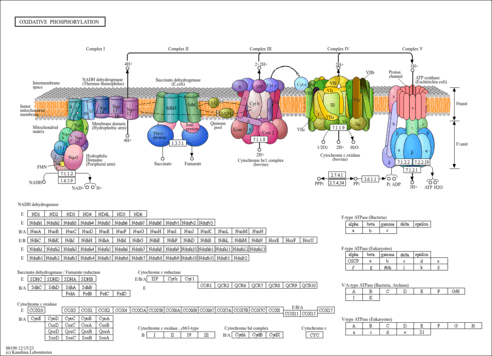
|
|---|
| Module |
|
|---|
| Organism |
Janthinobacterium sp. Marseille [GN: mms]
|
|---|
| Gene |
| mma_1506 | sdhC; succinate dehydrogenase cytochrome b-556 subunit [KO:K00241] |
| mma_1507 | sdhD; succinate dehydrogenase hydrophobic membrane anchor protein [KO:K00242] |
| mma_2637 | cyoD; cytochrome o ubiquinol oxidase operon protein [KO:K02300] |
| mma_3150 | cox11; cytochrome c oxidase subunit XI assembly protein [KO:K02258] |
| mma_3626 | atpC; F-type H+-transporting ATPase epsilon chain [KO:K02114] |
|
|---|
| Compound |
|
|---|
| Reference |
|
|---|
| Authors |
Sazanov LA, Hinchliffe P. |
|---|
| Title |
Structure of the hydrophilic domain of respiratory complex I from Thermus thermophilus. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Hinchliffe P, Carroll J, Sazanov LA. |
|---|
| Title |
Identification of a novel subunit of respiratory complex I from Thermus thermophilus. |
|---|
| Journal |
|
|---|
| KO pathway |
|
|---|