| Entry |
|
|---|
| Name |
Glutathione metabolism - Rattus norvegicus (rat) |
|---|
| Class |
Metabolism; Metabolism of other amino acids
|
|---|
| Pathway map |
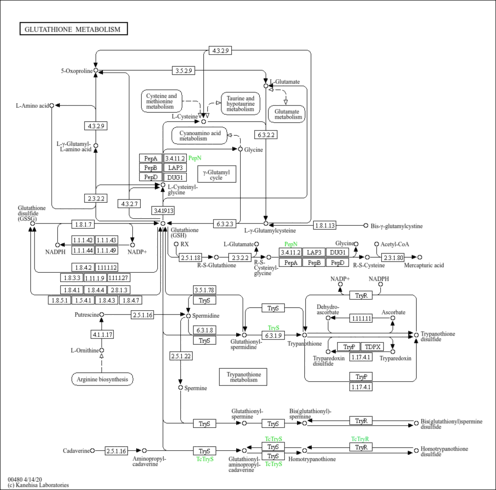
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Rattus norvegicus (rat) [GN: rno]
|
|---|
| Gene |
| 29328 | Gpx4; phospholipid hydroperoxide glutathione peroxidase isoform A precursor [KO:K05361] [EC:1.11.1.12] |
| 293656 | Gstp3; Glutathione S-transferase P-like isoform 2 [KO:K23790] |
| 29739 | Gclm; glutamate--cysteine ligase regulatory subunit [KO:K11205] |
| 360994 | Chac2; putative glutathione-specific gamma-glutamylcyclotransferase 2 [KO:K07232] [EC:4.3.2.7] |
|
|---|
| Compound |
| C00669 | gamma-L-Glutamyl-L-cysteine |
| C03170 | Trypanothione disulfide |
| C03646 | Bis-gamma-glutamylcystine |
| C03740 | (5-L-Glutamyl)-L-amino acid |
| C05726 | S-Substituted L-cysteine |
| C05727 | S-Substituted N-acetyl-L-cysteine |
| C16563 | Bis(glutathionyl)spermine |
| C16564 | Bis(glutathionyl)spermine disulfide |
| C16566 | Glutathionylaminopropylcadaverine |
| C16568 | Homotrypanothione disulfide |
|
|---|
| Reference |
|
|---|
| Authors |
Josch C, Klotz LO, Sies H. |
|---|
| Title |
Identification of cytosolic leucyl aminopeptidase (EC 3.4.11.1) as the major cysteinylglycine-hydrolysing activity in rat liver. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Chu L, Lai Y, Xu X, Eddy S, Yang S, Song L, Kolodrubetz D. |
|---|
| Title |
A 52-kDa leucyl aminopeptidase from treponema denticola is a cysteinylglycinase that mediates the second step of glutathione metabolism. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Cappiello M, Lazzarotti A, Buono F, Scaloni A, D'Ambrosio C, Amodeo P, Mendez BL, Pelosi P, Del Corso A, Mura U. |
|---|
| Title |
New role for leucyl aminopeptidase in glutathione turnover. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Suzuki H, Kamatani S, Kim ES, Kumagai H. |
|---|
| Title |
Aminopeptidases A, B, and N and dipeptidase D are the four cysteinylglycinases of Escherichia coli K-12. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Oza SL, Shaw MP, Wyllie S, Fairlamb AH. |
|---|
| Title |
Trypanothione biosynthesis in Leishmania major. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Oza SL, Ariyanayagam MR, Fairlamb AH. |
|---|
| Title |
Characterization of recombinant glutathionylspermidine synthetase/amidase from Crithidia fasciculata. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Tetaud E, Manai F, Barrett MP, Nadeau K, Walsh CT, Fairlamb AH. |
|---|
| Title |
Cloning and characterization of the two enzymes responsible for trypanothione biosynthesis in Crithidia fasciculata. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Ariyanayagam MR, Oza SL, Mehlert A, Fairlamb AH. |
|---|
| Title |
Bis(glutathionyl)spermine and other novel trypanothione analogues in Trypanosoma cruzi. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Oza SL, Ariyanayagam MR, Aitcheson N, Fairlamb AH. |
|---|
| Title |
Properties of trypanothione synthetase from Trypanosoma brucei. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Oza SL, Tetaud E, Ariyanayagam MR, Warnon SS, Fairlamb AH. |
|---|
| Title |
A single enzyme catalyses formation of Trypanothione from glutathione and spermidine in Trypanosoma cruzi. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Comini M, Menge U, Wissing J, Flohe L. |
|---|
| Title |
Trypanothione synthesis in crithidia revisited. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Hunter KJ, Le Quesne SA, Fairlamb AH. |
|---|
| Title |
Identification and biosynthesis of N1,N9-bis(glutathionyl)aminopropylcadaverine (homotrypanothione) in Trypanosoma cruzi. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Krauth-Siegel RL, Meiering SK, Schmidt H. |
|---|
| Title |
The parasite-specific trypanothione metabolism of trypanosoma and leishmania. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Krauth-Siegel RL, Comini MA. |
|---|
| Title |
Redox control in trypanosomatids, parasitic protozoa with trypanothione-based thiol metabolism. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Krauth-Siegel RL, Ludemann H. |
|---|
| Title |
Reduction of dehydroascorbate by trypanothione. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Dormeyer M, Reckenfelderbaumer N, Ludemann H, Krauth-Siegel RL. |
|---|
| Title |
Trypanothione-dependent synthesis of deoxyribonucleotides by Trypanosoma brucei ribonucleotide reductase. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Schmidt H, Krauth-Siegel RL. |
|---|
| Title |
Functional and physicochemical characterization of the thioredoxin system in Trypanosoma brucei. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Tetaud E, Fairlamb AH. |
|---|
| Title |
Cloning, expression and reconstitution of the trypanothione-dependent peroxidase system of Crithidia fasciculata. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Castro H, Sousa C, Santos M, Cordeiro-da-Silva A, Flohe L, Tomas AM. |
|---|
| Title |
Complementary antioxidant defense by cytoplasmic and mitochondrial peroxiredoxins in Leishmania infantum. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Wilkinson SR, Temperton NJ, Mondragon A, Kelly JM. |
|---|
| Title |
Distinct mitochondrial and cytosolic enzymes mediate trypanothione-dependent peroxide metabolism in Trypanosoma cruzi. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Konig J, Fairlamb AH. |
|---|
| Title |
A comparative study of type I and type II tryparedoxin peroxidases in Leishmania major. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Hillebrand H, Schmidt A, Krauth-Siegel RL. |
|---|
| Title |
A second class of peroxidases linked to the trypanothione metabolism. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Soksawatmaekhin W, Kuraishi A, Sakata K, Kashiwagi K, Igarashi K. |
|---|
| Title |
Excretion and uptake of cadaverine by CadB and its physiological functions in Escherichia coli. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Pegg AE, Shuttleworth K, Hibasami H. |
|---|
| Title |
Specificity of mammalian spermidine synthase and spermine synthase. |
|---|
| Journal |
|
|---|
Related
pathway |
| rno00250 | Alanine, aspartate and glutamate metabolism |
| rno00270 | Cysteine and methionine metabolism |
| rno00430 | Taurine and hypotaurine metabolism |
|
|---|
| KO pathway |
|
|---|