PROSITE is a database of protein families and domains. It consists of biologically significant sites, patterns and profiles that help to reliably identify to which known protein family (if any) a new sequence belongs.
The generalized profiles developed by Bucher, P. and Bairoch A. is very sensitive method to find motifs in a query sequence. Profiles created from various protein domains are incorporated into the PROSITE library. Here a program called Findprofile written in ICR, Kyoto University has been used to search with a sequence against profile database. This program uses dynamic programming algorhism to find out the best alignment between a query sequence and each profile entry in PROSITE dabase. Only the profiles which have scores greater than given threshould are shown.
When a profile is found, wherever available, series of three dimensional structures are retrieved from PDB database, which share the motif and you can examine them using RasMol program the same as the result of pattern database search.
ID ACP_DOMAIN; MATRIX. AC PS50075; DT NOV-1997 (CREATED); NOV-1997 (DATA UPDATE); JUL-1998 (INF UPDATE). DE Acyl carrier protein phosphopantetheine domain profile. MA /GENERAL_SPEC: ALPHABET='ABCDEFGHIKLMNPQRSTVWYZ'; LENGTH=71; MA /DISJOINT: DEFINITION=PROTECT; N1=6; N2=66; MA /NORMALIZATION: MODE=1; FUNCTION=LINEAR; R1=2.3; R2=.02281121; TEXT='NScore'; MA /CUT_OFF: LEVEL=0; SCORE=271; N_SCORE=8.5; MODE=1; MA /CUT_OFF: LEVEL=-1; SCORE=184; N_SCORE=6.5; MODE=1;In this example two "levels" of cut-off score are annotated.
Positive Score ad increases the cut-off threshould and vice versa.
You can search your query sequnce against a profile library defined by a user which contains
either single or multiple profile data in PROSITE format. Check the "User-defined
Profile Library" box in the motif library list and provide a file name containing the
profile.
The user defined profile library may be a subset of original PROSITE database or one generated
from multiple sequence alignment data.
Common sequence patterns such as "C-kinase phosphorylation site" or " N-glycosylation site" (which are assigned as "/SKIP-FLAG=TRUE" on the CC line in the database) are ignored automatically.
If any profiles are found in the database ProfileFinder then looks up the Protein Data Bank
entries which share the same motif annotated in PROSITE dabase on 3D lines.
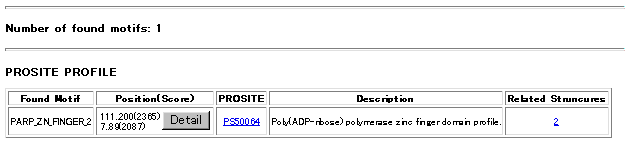
Under Related Structures column you can find the number of those structures.
A list of those entries, ID numbers of PDB together with brief description of the proteins is shown by clicking the cell. From the list you can either jump into DBGET to look at the PDB entry more precisely,
or see the position and the structre of the profile motif on the 3D structure.
Under Position (Score) column of the table the position (start and end sequence numbers) and the raw score values of found motifs are listed. Click Detail bottun to see actual positions of the motif along the query sequence . A Consensus sequence is taken from each database entry. Aligned with this consensus sequence a fragment of query sequence (upper one) is shown with the score. Actual position on the query sequence is also shown. (red residue letters)
Below a sample output table of the search is shown. Click the images to get detailed results discribed above.

|